Session VI - Avian cytogenetics
L7
Evolution of the avian genome as revealed by molecular cytogenetics
Martin Völker (1), Benjamin M. Skinner (1), Helen G. Tempest (2), Darren K. Griffin (1)
1. Department of Biosciences, University of Kent, Canterbury, Kent CT2 7 NJ (UK)
2. Department of Medical Genetics, University of Calgary, Calgary T2N 4N1 (Canada)
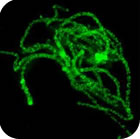
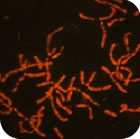
In avian genome evolution, the diploid number of 2n≈80 has remained remarkably constant with 63% of birds having 2n=74-86. The most studied species is the chicken and molecular cytogenetic probes generated by us from this species have been used to understand further the evolution of the avian genome. The ancestral karyotype is very similar to that of the chicken, with chicken chromosomes 1, 2, 3, 4q, 5, 6, 7, 8, 9, 4p and Z representing the ancestral avian chromosomes 1-10+Z. Avian evolution occurred primarily in three stages: divergence of the Paleognathae (emu, ostrich etc.); divergence of the Galloanserae (chicken, goose etc.); and divergence of the higher "land" and higher "water" birds. Other than sex chromosome differentiation in the first divergence there are no specific changes associated with any of these evolutionary milestones although certain groups (e.g. the Falconiformes and the Psittaciformes) have undergone multiple fusions (and some fissions). Most changes seem to involve chromosomes 1, 2, 4, 6-9, 10 and Z and there are several convergent events. Of these, the most puzzling involves chromosomes 4 and 10, which appear to have undergone multiple fissions and fusions throughout evolution. The use of microarrays and detailed cytogenetic maps is shedding further light on avian genome organization and evolution.
O13
Nuclear (genome) organisation and comparative genomics in birds
Benjamin M. Skinner (1), Martin Völker (1), Gothami L. Fonseka (1), Michael Ellis (2), Darren K. Griffin (1)
1. Department of Biosciences, University of Kent, Canterbury, Kent, CT2 7NJ (UK)
2. Digital Scientific Ltd, Sheraton House, Castle Park, Cambridge, CB3 0AX (UK)
Nuclear, or genome organisation - the position of chromosome territories within the interphase nucleus - is recognised as playing an important role in development, disease and evolution. Two models for genome organisation suggest that chromosomes can be organised either by chromosome size or by gene density. As a key model species and agricultural animal, the chicken (Gallus gallus) has the best studied avian genome organisation; the macrochromosomes tend to reside at the periphery of the nucleus, while the microchromosomes are more internal. The precise nuclear positioning of the smaller macrochromosomes and microchromosomes in chicken has not however been determined, nor has the model it fits more closely.
Here, nuclear addresses of macro- and microchromosome territories in chicken were determined by measuring the positions of BAC clones within fibroblast nuclei. The areas of metaphase chromosomes were measured, and gene densities were calculated. The chromosomes were ordered by size and by gene density to determine which model chicken follows more closely; results suggest no specific preference for either model. This study was also carried out in turkey (Meleagris gallopovo) and duck (Anas platyrhynchos), using the same BACs applied to chicken, to determine whether chromosomes involved in fusions or fissions occupy different positions in their ancestral and derived states.